The free BEHST software package efficiently associates functional enriched Gene Ontology terms to input genomic regions
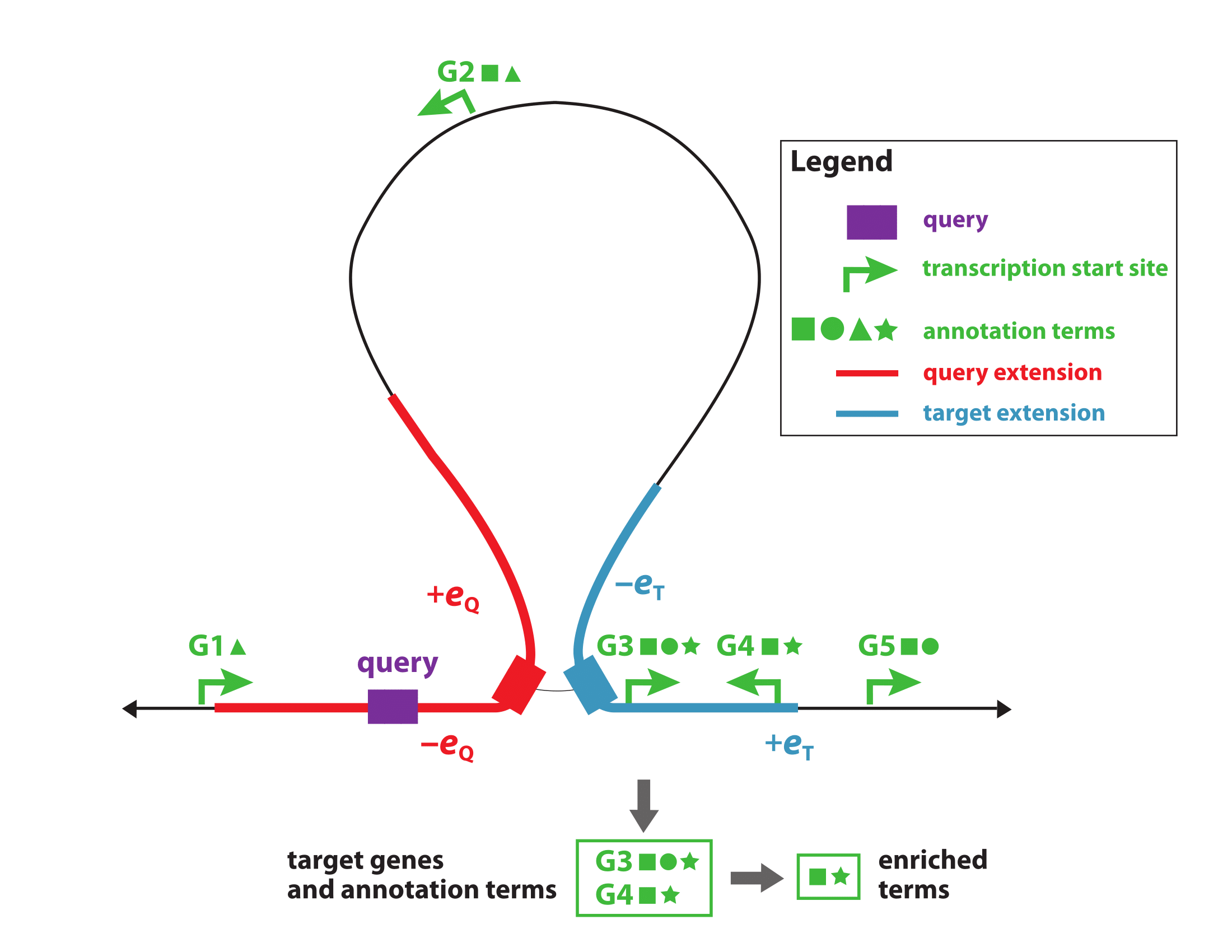
BEHST reads a dataset of genomic regions, and intersects them with the chromatin interactions available in the Hi-C dataset (Rao et al, Cell, 2014). Of these genomic regions, BEHST selects those that are present in the regulatory regions of genes a dataset of principal isoform annotations. We defined these cis-regulatory regions upon the position of their nearest transcription start site of the genes' principal transcripts, plus an upstream and downstream extension. Afterwards, BEHST selects the genes of the resulting partner loci found in gene regulatory regions, and inserts them into g:Profiler. BEHST, finally, produces the list of the most significant Gene Ontology terms detected by g:Profiler.
Installation
BEHST can run on any Linux and Mac computers. You can find the installation instructions on BEHST Bitbucket webpage.
Support
For support of BEHST, please write to the BEHST-users mailing list, rather than writing the authors directly. Using the mailing list will get your question answered more quickly. It also allows us to pool knowledge and reduce getting the same inquiries over and over. Questions sent to the mailing list will
If you do not want to read discussions about other people's use of BEHST, but would like to hear about new releases and other important information, please subscribe to the BEHST-announce mailing list mailing list.
Source code
BEHST code repository on Bitbucket
Credits
BEHST was developed by Davide Chicco, Haixin Sarah Bi, Juri Reimand, and Michael M. Hoffman at the Hoffman Lab of the Princess Margaret Cancer Centre (Toronto, Ontario, Canada). This website was developed by Abdellatif Daghrach, Dustin Hu, and Kevin Chan of the Princess Margaret Genomics Centre (Toronto, Ontario, Canada).